The design of new proteins and enzymes remains one of the great challenges in biochemistry and tests our fundamental understanding of both the nature of protein as a material and the principles of enzymatic catalysis. Unlocking the exceptionally diverse and powerful array of chemistries exhibited by natural enzymes promises routes to new drugs, therapies and green industrial processes. Most approaches to this end have focused on modifying natural enzymes to impart new or altered catalytic function. The problems that often hinder the re-engineering of naturally evolved proteins and enzymes are due to the layers of complexity that nature incorporates through natural selection into a protein’s complex 3D structure.
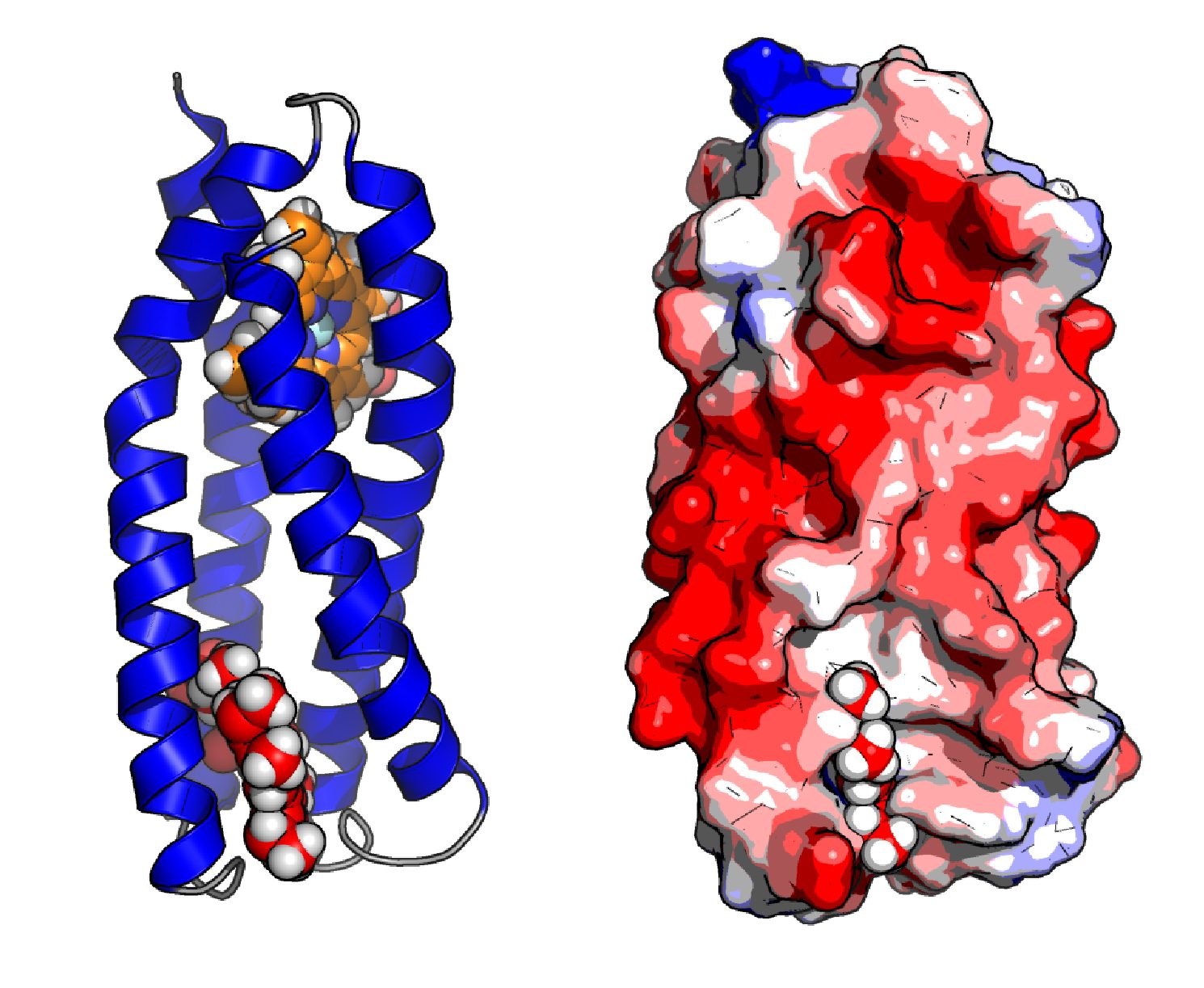
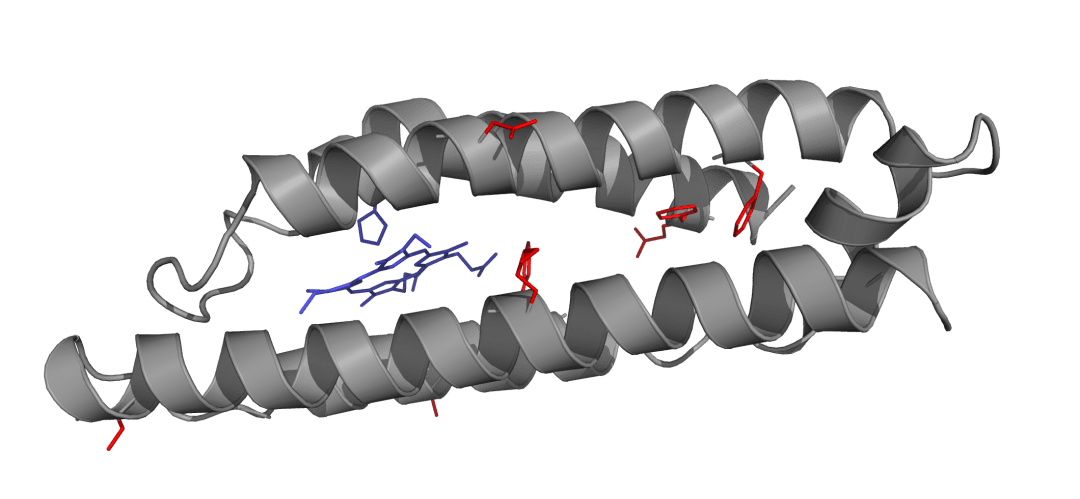
Simplified manmade protein scaffolds offer a means to avoid such complexity, learn the principles guiding functional protein assembly and render the modular assembly of enzymatic function a tangible reality. This approach is illustrated through the assembly of artificial oxygen binding proteins that reproduce the function of natural proteins such as myoglobin in simple heme-binding 4-helix bundles untouched by natural selection (Koder & Anderson et al., Nature, 2009) The tractable design process that we employ resolves the roles of individual amino acids with their function and opens the door to the powerful oxygenic catalysis common to heme-containing enzymes.
In the Anderson laboratory, we use this simple protein design approach to construct artificial oxidoreductase enzymes that integrate functional elements common to natural redox enzymes – e.g. electron/proton transfer, ligand/substrate binding and light harvesting – in a discrete manmade protein that is wholly fabricated within a living organism (Anderson et al. Chemical Science, 2014; Watkins et al. BBA-Bioenergetics, 2016).